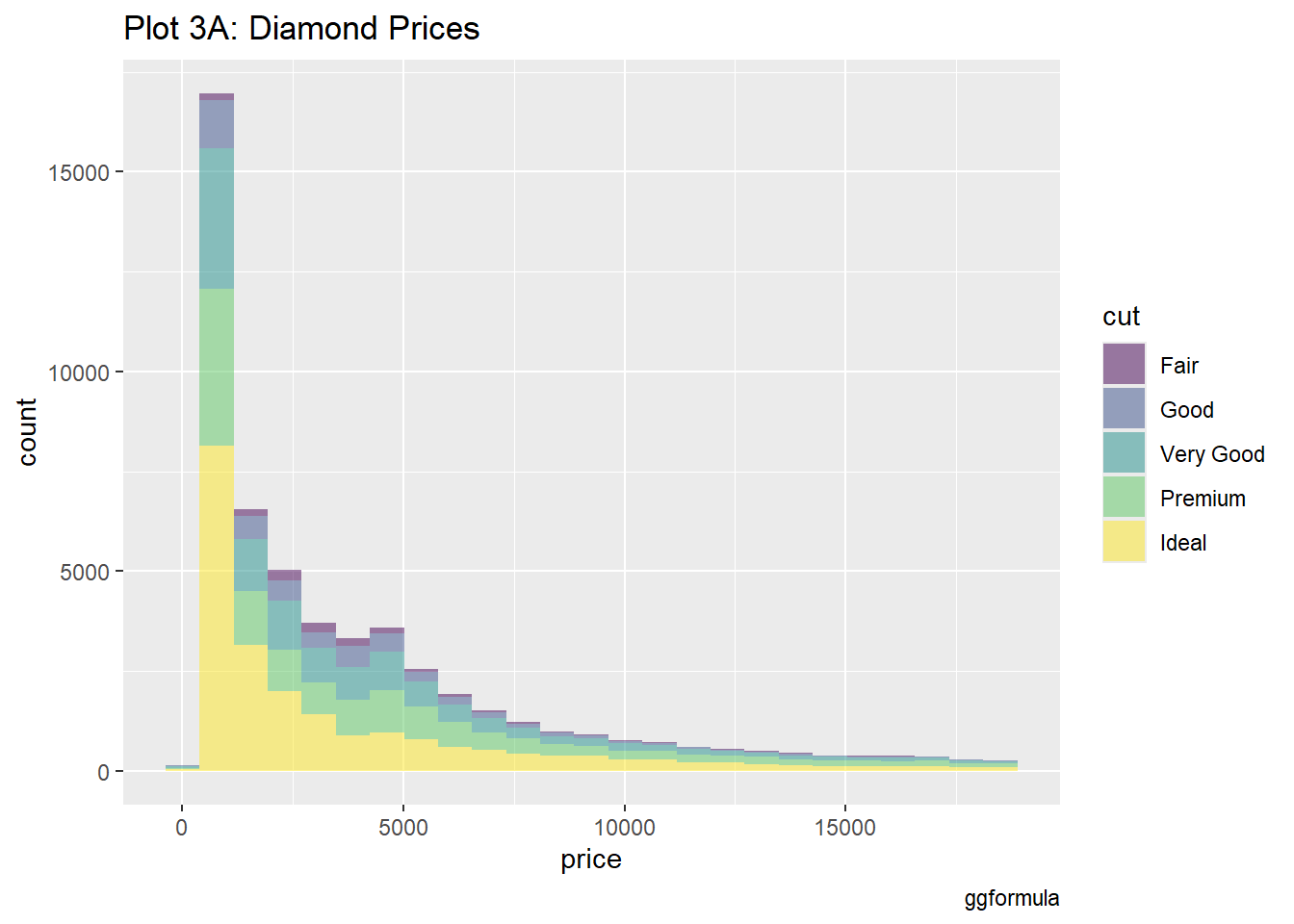
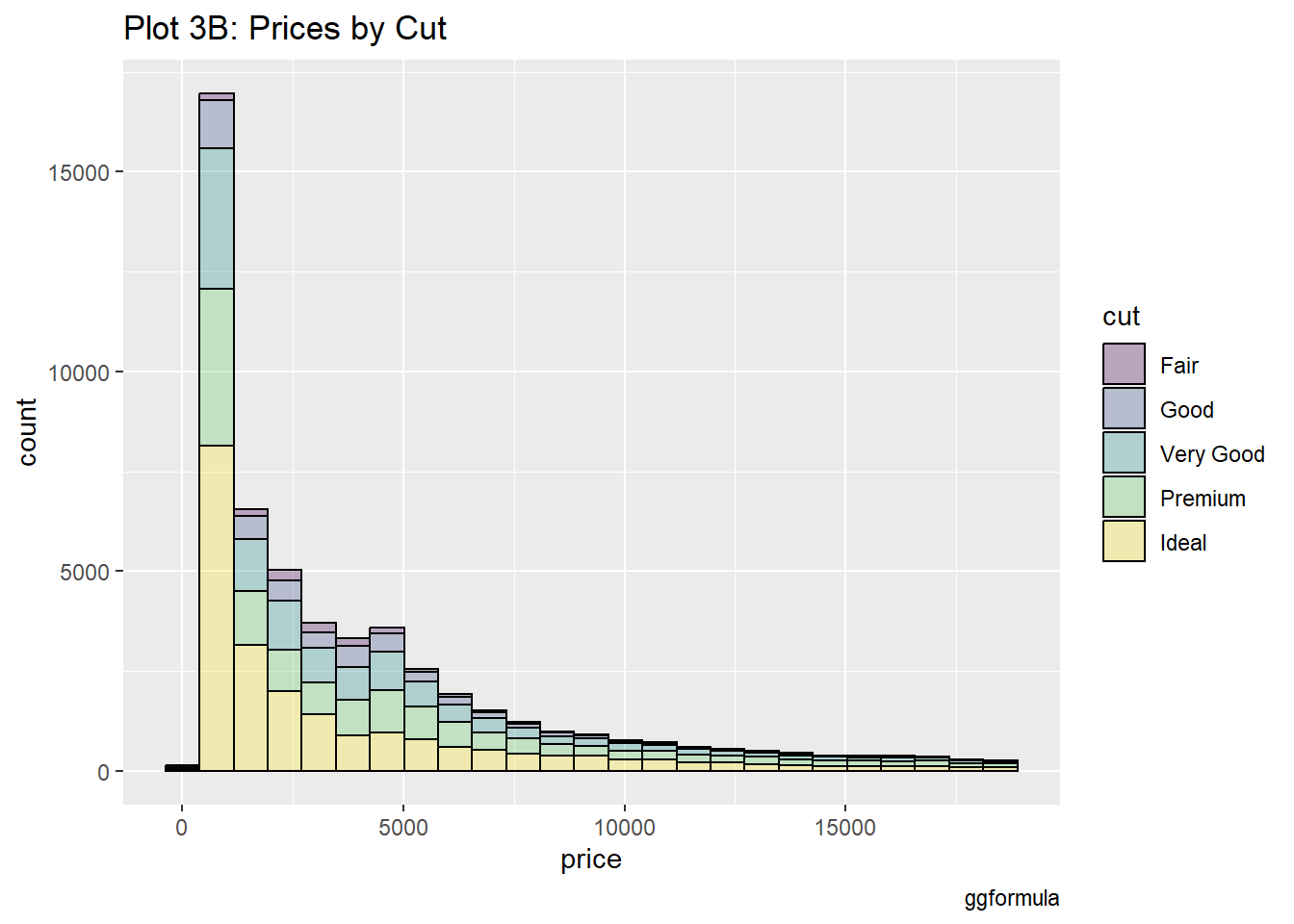
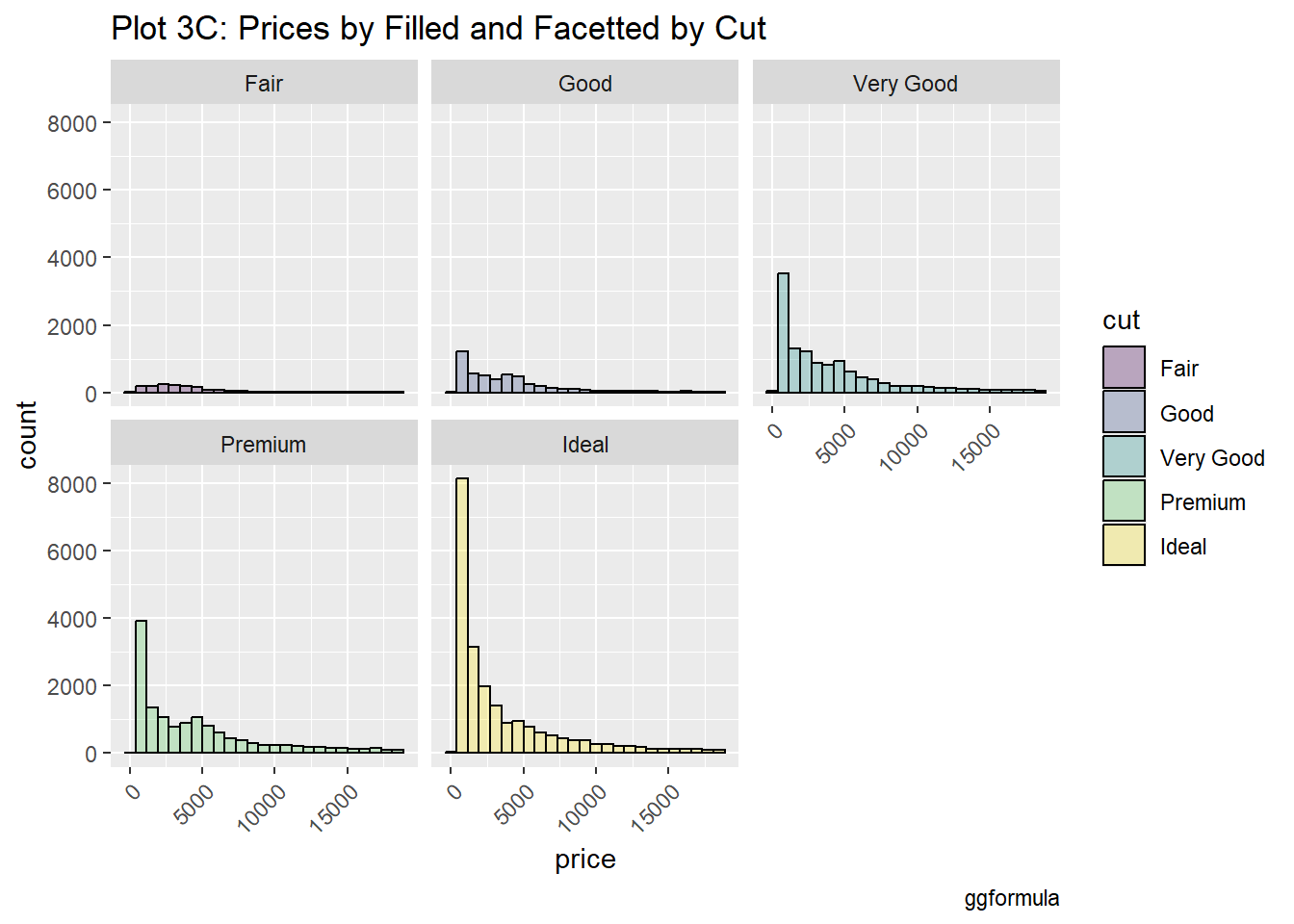
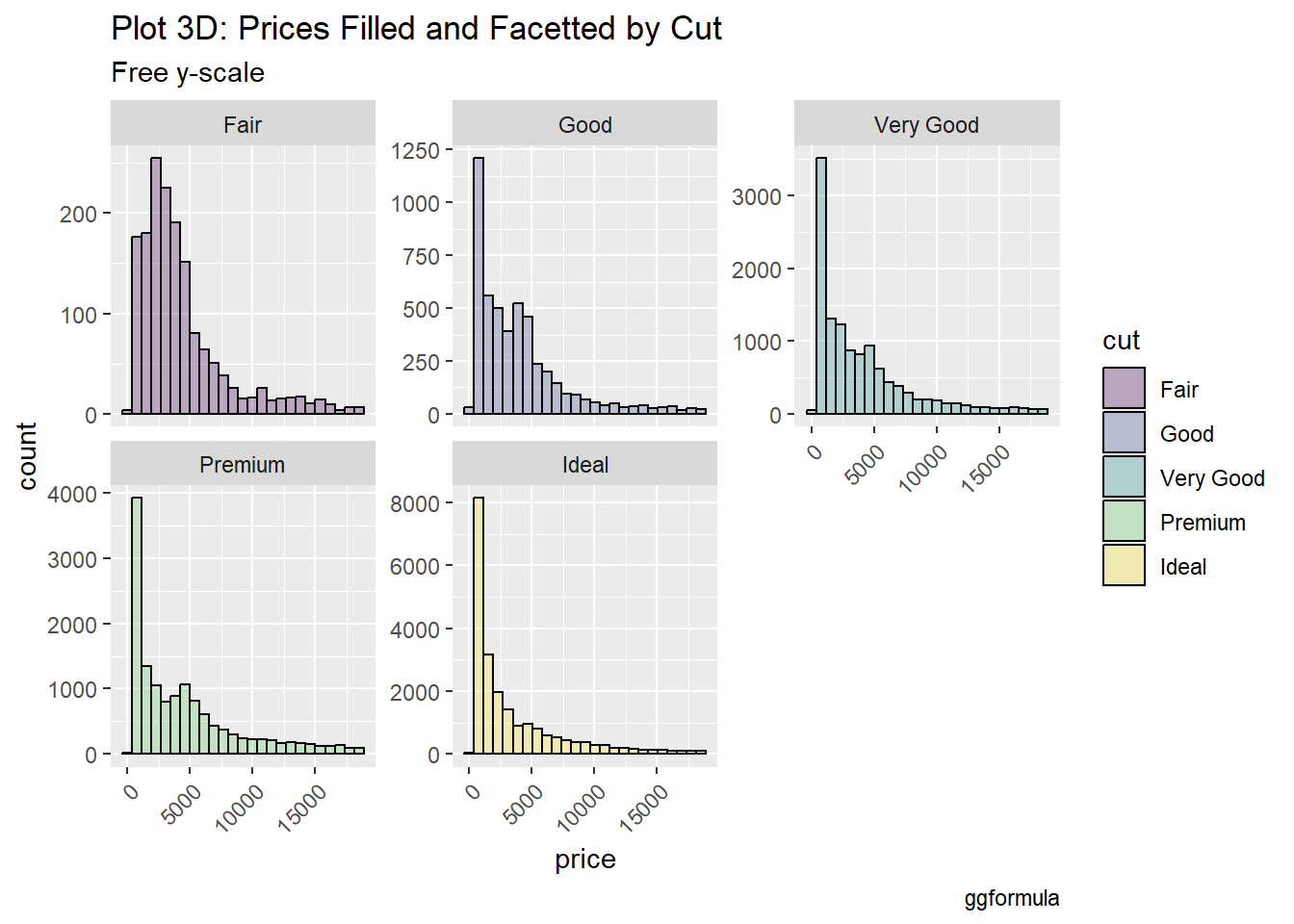
#|label: setup
library(tidyverse)── Attaching core tidyverse packages ──────────────────────── tidyverse 2.0.0 ──
✔ dplyr 1.1.4 ✔ readr 2.1.5
✔ forcats 1.0.0 ✔ stringr 1.5.1
✔ ggplot2 3.5.1 ✔ tibble 3.2.1
✔ lubridate 1.9.3 ✔ tidyr 1.3.1
✔ purrr 1.0.2
── Conflicts ────────────────────────────────────────── tidyverse_conflicts() ──
✖ dplyr::filter() masks stats::filter()
✖ dplyr::lag() masks stats::lag()
ℹ Use the conflicted package (<http://conflicted.r-lib.org/>) to force all conflicts to become errorslibrary(mosaic)Registered S3 method overwritten by 'mosaic':
method from
fortify.SpatialPolygonsDataFrame ggplot2
The 'mosaic' package masks several functions from core packages in order to add
additional features. The original behavior of these functions should not be affected by this.
Attaching package: 'mosaic'
The following object is masked from 'package:Matrix':
mean
The following objects are masked from 'package:dplyr':
count, do, tally
The following object is masked from 'package:purrr':
cross
The following object is masked from 'package:ggplot2':
stat
The following objects are masked from 'package:stats':
binom.test, cor, cor.test, cov, fivenum, IQR, median, prop.test,
quantile, sd, t.test, var
The following objects are masked from 'package:base':
max, mean, min, prod, range, sample, sumlibrary(ggformula)
library(skimr)
Attaching package: 'skimr'
The following object is masked from 'package:mosaic':
n_missinglibrary(crosstable)
Attaching package: 'crosstable'
The following object is masked from 'package:purrr':
compact